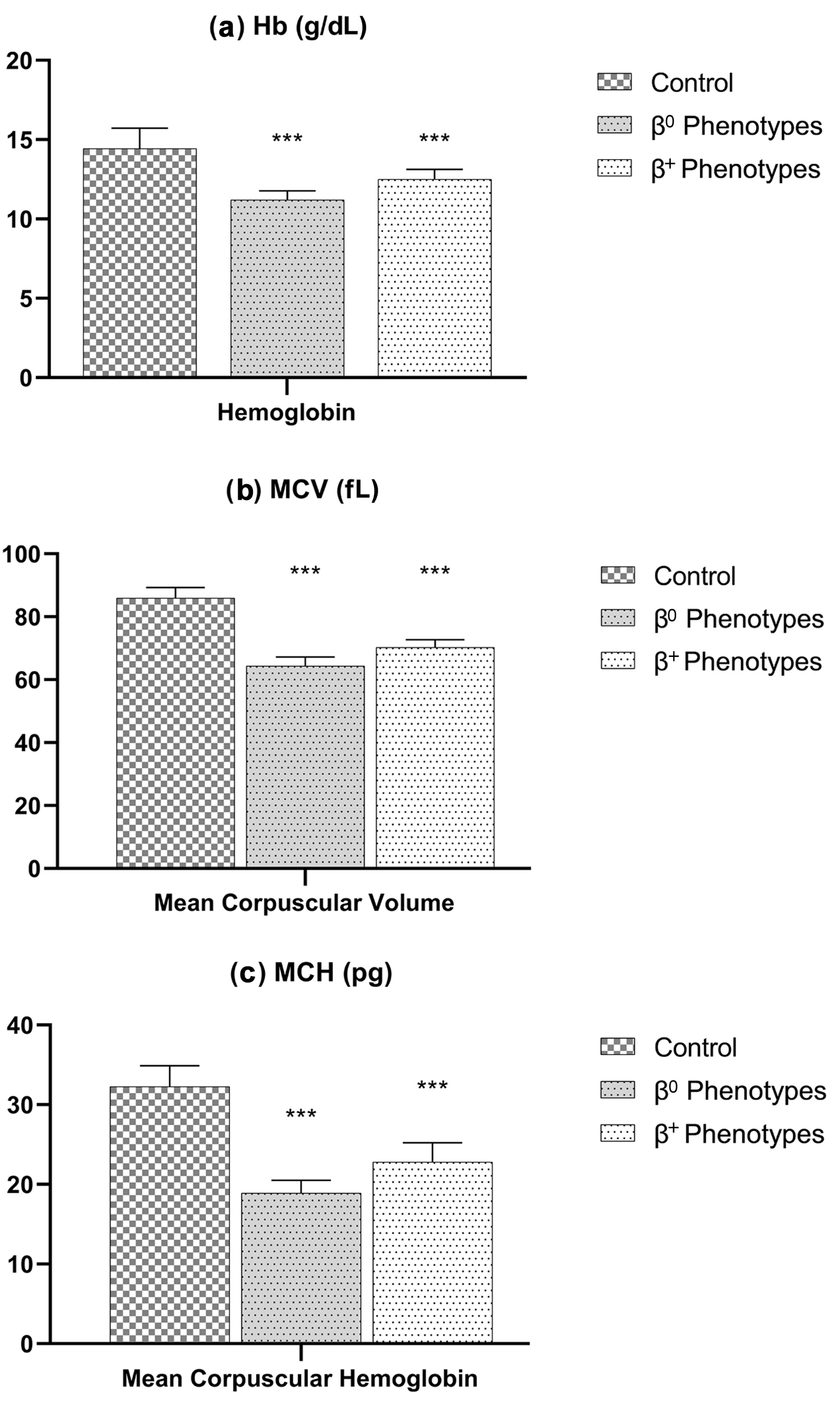
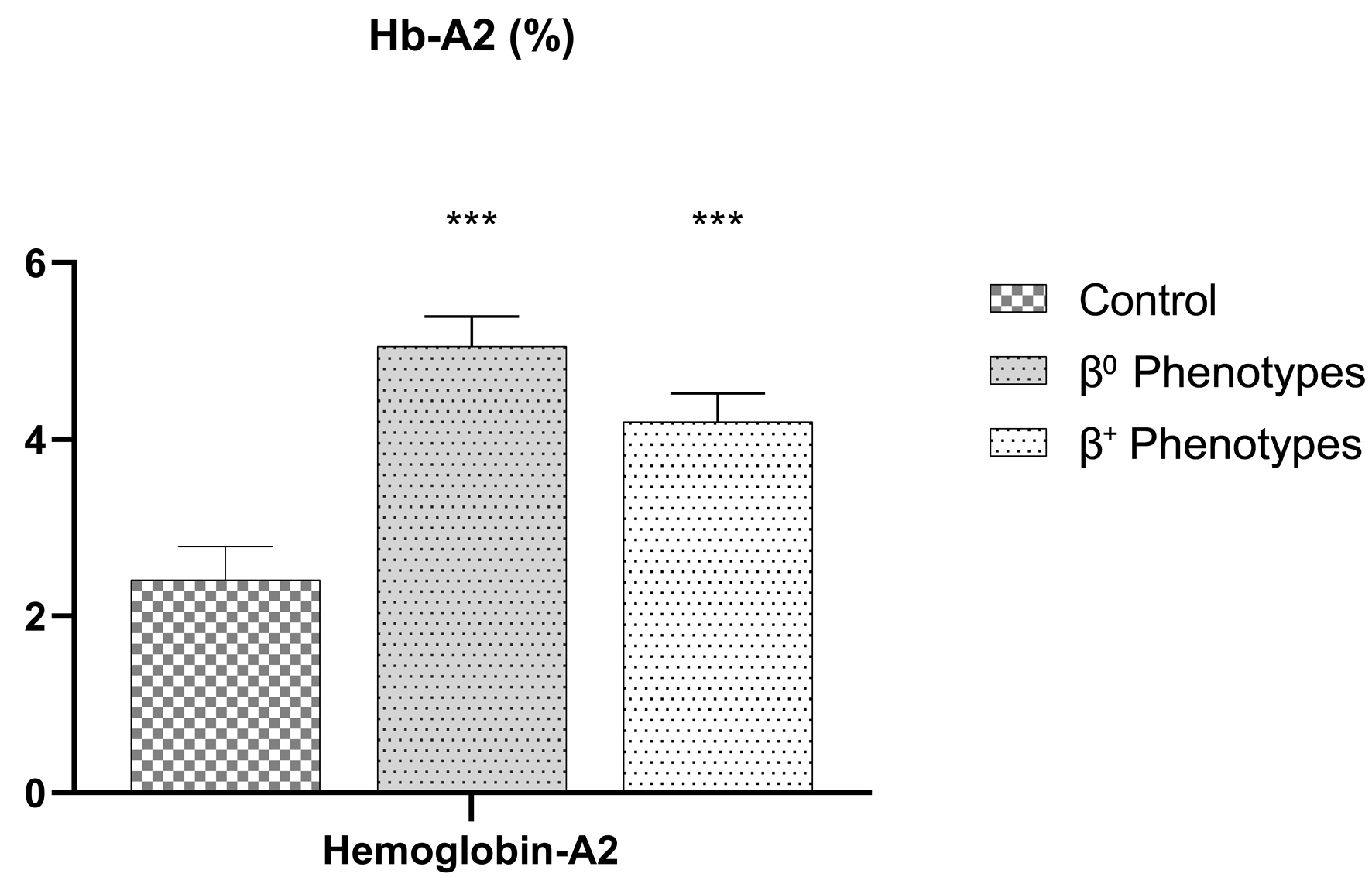
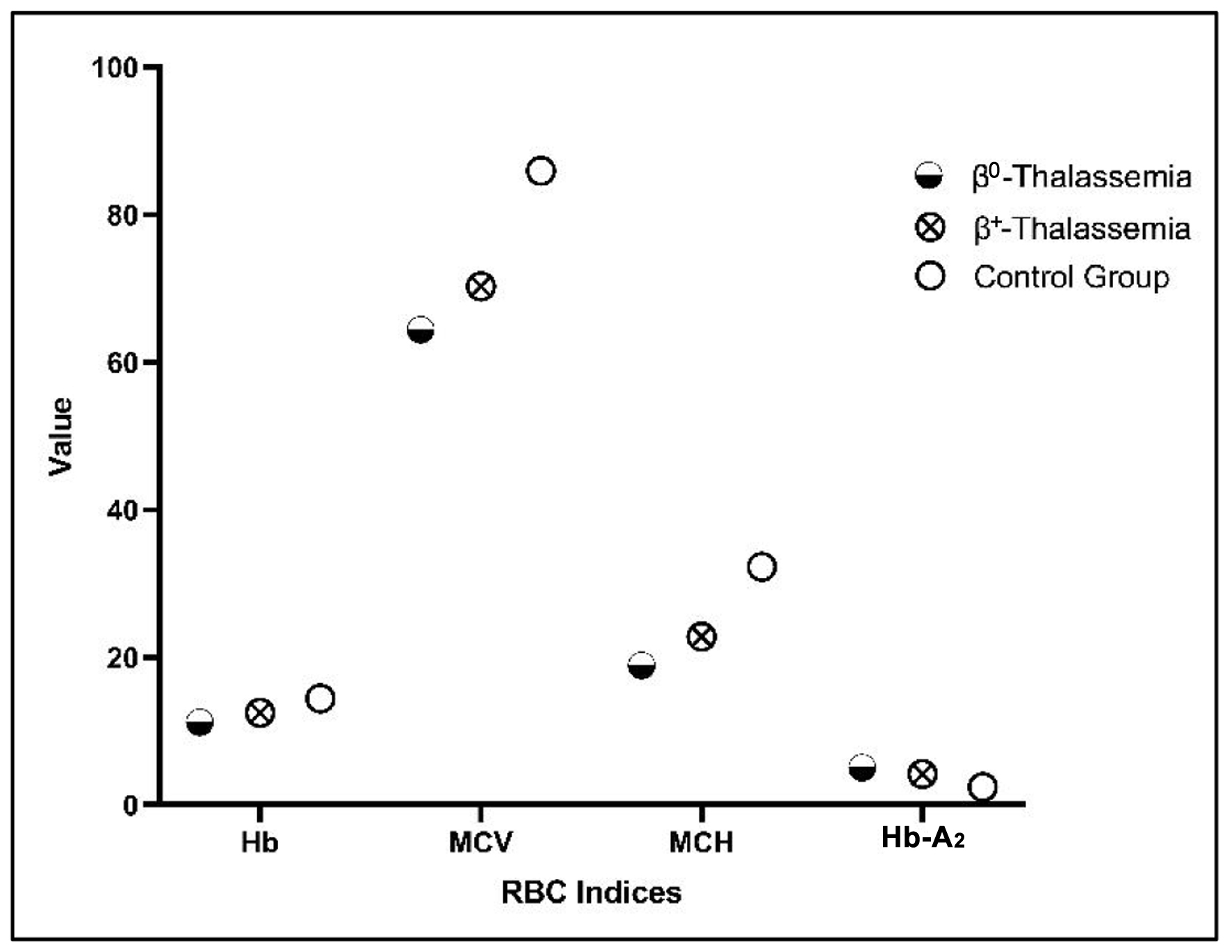
| β0 phenotypes (N = 64) (44.1%) |
|
|
|
|
|
|
| IVS 2.1 [G>A] |
HBB: c.315+1G>A |
RNA processing/splice junction |
β0 |
Mediterranean, US Blacks |
21 |
32.8 |
| IVS 1.1 [G>A] |
HBB: c.92+1G>A |
RNA processing/splice junction |
β0 |
Mediterranean |
18 |
28.1 |
| Codon 8 [-AA] |
HBB: c.25_26delAA |
RNA translation/frameshift |
β0 |
Mediterranean |
8 |
12.5 |
| Codon 5 [-CT] |
HBB: c.17_18delCT |
RNA translation/frameshift |
β0 |
Mediterranean |
7 |
10.9 |
| Codon 39 [C>T] |
HBB: c.118C>T |
RNA translation/nonsense codons |
β0 |
Mediterranean |
7 |
10.9 |
| Codon 6 [-A] |
HBB: c.20delA |
RNA translation/frameshift |
β0 |
Mediterranean, US Blacks |
1 |
1.6 |
| IVS 1.130 [G>C] |
HBB: c.93-1G>C |
RNA processing/splice junction |
β0 |
Italian, Japanese, UAE |
1 |
1.6 |
| Codon 44 [-C] |
HBB: c.135delC |
RNA translation/frameshift |
β0 |
Kurdish |
1 |
1.6 |
| β+ phenotypes (N = 81) (55.9%) |
|
|
|
|
|
|
| IVS 1.6 [T>C] |
HBB: c.92+6T>C |
RNA processing/consensus splice sites |
β+ |
Mediterranean |
28 |
34.6 |
| IVS 1.110 [G>A] |
HBB: c.93-21G>A |
RNA processing/cryptic splice sites |
β+ |
Mediterranean |
21 |
25.9 |
| IVS 2.745 [C>G] |
HBB: c.316-106C>G |
RNA processing/cryptic splice sites |
β+ |
Mediterranean |
16 |
19.8 |
| IVS 1.5 [G>C] |
HBB: c.92+5G>C |
RNA processing/consensus splice sites |
β+ |
Asian Indian, SE Asian |
5 |
6.2 |
| -30 [T>A] |
HBB: c.-80T>A |
Transcriptional variants/promoter regulatory elements |
β+ |
Mediterranean, Bulgarian |
4 |
4.9 |
| -87 [C>G] |
HBB: c.-137C>G |
Transcriptional variants/promoter regulatory elements |
β++ |
Mediterranean |
3 |
3.7 |
| -101 [C>T] |
HBB: c.-151C>T |
Transcriptional variants/promoter regulatory elements |
β++ (silent) |
Mediterranean |
3 |
3.7 |
| IVS 2.848 [C>A] |
HBB: c.316-3C>A |
RNA processing/consensus splice sites |
β+ |
UB Blacks, Egyptian, Iranian |
1 |
1.2 |